RStudio Conference 2019 takes place in January 2019, and this week RStudio put out a call for contributed talks and e-posters. Though I was eager to browse previous years’ abstracts for inspiration, I couldn’t find them all in one place, and so I decided to use one of my favourite R packages, rvest, to do some web scraping to grab the content.
My main aim was to find all of the abstracts for the contributed talks only from 2018.
As this ended up being an unusually long blog post, here’s a table containing links to the videos and abstract for contributed talks. A detailed walk through of the code used to create it can be found below.
As ever, give me a shout on Twitter if you have any comments or questions!
Here’s a walkthrough of the code, and the full list of abstracts.
Getting Started
I found last year’s conference schedule here, which seemed as good a place as any to start. First, I loaded the rvest package and used read_html() (from xml2 which is automatically loaded when you load rvest) to pull the html from the page.
library(rvest)
my_html <- read_html("https://beta.rstudioconnect.com/content/3105/")
my_html## {xml_document}
## <html xmlns="http://www.w3.org/1999/xhtml">
## [1] <head>\n<meta charset="utf-8">\n<meta http-equiv="Content-Type" cont ...
## [2] <body>\n\n<style type="text/css">\n.main-container {\n max-width: 9 ...The my_html object is an xml document, and to do anything useful with it, I need to work out which components I need. I could do this manually, but it’s much simpler to use Selector Gadget to help me.
Selector Gadget
Selector Gadget is a tool which takes a lot of effort out of webscraping. For an in-depth description, check out this rvest vignette. In short, once Selector Gadget is loaded, I click on any unselected components to pick which ones I need (in green), look at what else is highlighted (in yellow) and then click on these to deselect them (now in red) and carry on selecting and/or deselecting components until only the ones I want are in green or yellow.
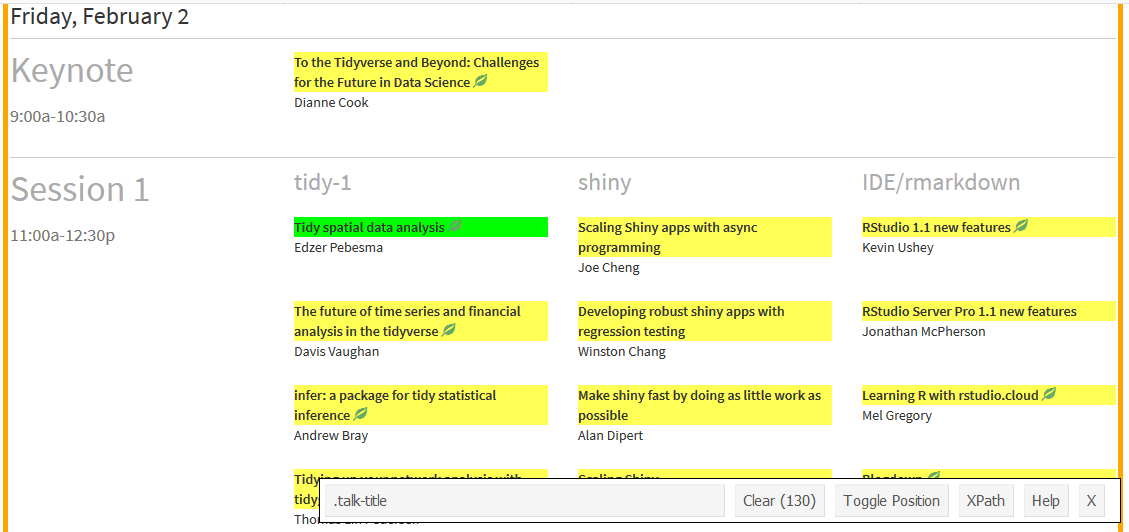
Then, I copy the text out of the box, in the case, “.talk-title”. This is the CSS selector needed to grab those components, and I supply this as the css argument to html_nodes(). As I just want the inner text, I pipe the result of this into html_text().
talks <- my_html %>%
html_nodes(".talk-title") %>%
html_text()
head(talks)## [1] "\nTo the Tidyverse and Beyond: Challenges for the Future in Data Science\n\n"
## [2] "\nTo the Tidyverse and Beyond: Challenges for the Future in Data Science\n\n"
## [3] "\nTidy spatial data analysis\n\n"
## [4] "Scaling Shiny apps with async programming"
## [5] "\nRStudio 1.1 new features\n\n"
## [6] "\nThe future of time series and financial analysis in the tidyverse\n\n"I now repeat this process to get the presenter names. Handily, the CSS selector for these is “.presenter”!
authors <- my_html %>%
html_nodes(".presenter") %>%
html_text()
head(authors)## [1] "Dianne Cook" "Dianne Cook" "Edzer Pebesma" "Joe Cheng"
## [5] "Kevin Ushey" "Davis Vaughan"Now I’ve got my talks and authors, I’m next going to combine them in a data_frame. Once this is done, I’m also going to tidy them up by removing all the newline characters (“”) from the titles.
library(dplyr)
library(stringr)
schedule <- data_frame(talks, authors) %>%
mutate(talks = str_remove_all(talks, "\n")) %>%
distinct()
head(schedule)## # A tibble: 6 x 2
## talks authors
## <chr> <chr>
## 1 To the Tidyverse and Beyond: Challenges for the Future in D~ Dianne Cook
## 2 Tidy spatial data analysis Edzer Pebe~
## 3 Scaling Shiny apps with async programming Joe Cheng
## 4 RStudio 1.1 new features Kevin Ushey
## 5 The future of time series and financial analysis in the tid~ Davis Vaug~
## 6 Developing robust shiny apps with regression testing Winston Ch~Great! So now what? Well, the point of this analysis is to look at the contributed talks, rather than those from invited speakers or RStudio staff, so in the next part of this post, I’m going to be using lots of dplyr joins to remove these!
Finding Only Contributed Talks
Let’s start of with identifying talks from invited speakers and RStudio staff.
I found this post on the RStudio blog which lists the invited speakers. I repeat the process from earlier using rvest and xml2 functions to scrape the list of invited speakers.
invited_speakers <- read_html("https://blog.rstudio.com/2017/07/12/join-us-at-rstudioconf-2018/") %>%
html_nodes("table:nth-child(6) td:nth-child(1)") %>%
html_text()
invited <- data_frame(authors = invited_speakers, Status = "Invited")
head(invited)## # A tibble: 6 x 2
## authors Status
## <chr> <chr>
## 1 Mara Averick Invited
## 2 Nick Carchedi Invited
## 3 Tanya Cashorali Invited
## 4 Eric Colson Invited
## 5 Sandra Griffith Invited
## 6 Aaron Horowitz InvitedNow I need to combine this back with my original table, schedule, so I use left_join() from dplyr to help me.
author_tbl <- left_join(schedule, invited, by = "authors")
head(author_tbl)## # A tibble: 6 x 3
## talks authors Status
## <chr> <chr> <chr>
## 1 To the Tidyverse and Beyond: Challenges for the Fut~ Dianne Cook <NA>
## 2 Tidy spatial data analysis Edzer Peb~ Invit~
## 3 Scaling Shiny apps with async programming Joe Cheng <NA>
## 4 RStudio 1.1 new features Kevin Ush~ <NA>
## 5 The future of time series and financial analysis in~ Davis Vaugh~ <NA>
## 6 Developing robust shiny apps with regression testing Winston Ch~ <NA>Something you may have noticed is that we have some duplicated rows in the table, and so I use distinct() to remove these.
author_tbl <- distinct(author_tbl)
head(author_tbl)## # A tibble: 6 x 3
## talks authors Status
## <chr> <chr> <chr>
## 1 To the Tidyverse and Beyond: Challenges for the Fut~ Dianne Cook <NA>
## 2 Tidy spatial data analysis Edzer Peb~ Invit~
## 3 Scaling Shiny apps with async programming Joe Cheng <NA>
## 4 RStudio 1.1 new features Kevin Ush~ <NA>
## 5 The future of time series and financial analysis in~ Davis Vaugh~ <NA>
## 6 Developing robust shiny apps with regression testing Winston Ch~ <NA>Great! Now I’ve remove the duplications, let’s take a look at how many rows have no value in the Status column. I can use sum() and is.na() to do this
sum(is.na(author_tbl$Status))## [1] 51I still have 51 rows with no value in the Status column, but I’m only expecting around 20. I need to remove talks from RStudio staff to get to my contributed talks.
I did the next bit manually, by searching for any of the individuals I hadn’t already heard of. It’s likely that there’s a quicker way to do this, using the web scraping techniques we’ve already looked at.
In no particular order…
rstudio_staff <- c("Joe Cheng", "Winston Chang", "Alan Dipert", "Sean Lopp",
"Kevin Ushey", "Jonathan McPherson", "Mel Gregory",
"Yihui Xie", "Max Kuhn", "Jenny Bryan", "Jim Hester",
"Joseph Rickert", "Jeff Allen", "Aron Atkins",
"Barbara Borges Ribeiro", "Nathan Stephens",
"Mine Cetinkaya-Rundel", "Aaron Berg", "Hadley Wickham",
"Edgar Ruiz", "Amanda Gadrow", "JJ Allaire", "Kevin Kuo",
"Javier Luraschi", "Michael Quinn", "Tareef Kawaf")
rstudio <- data_frame(authors = rstudio_staff, Status = "RStudio")
head(rstudio)## # A tibble: 6 x 2
## authors Status
## <chr> <chr>
## 1 Joe Cheng RStudio
## 2 Winston Chang RStudio
## 3 Alan Dipert RStudio
## 4 Sean Lopp RStudio
## 5 Kevin Ushey RStudio
## 6 Jonathan McPherson RStudioOK, so this is where it gets a little complicated, so please bear with me.
What I need to do next is combine my list of RStudio people with the schedule data_frame. However, I can’t just do a left_join:
left_join(author_tbl, rstudio, by = "authors") %>%
head()## # A tibble: 6 x 4
## talks authors Status.x Status.y
## <chr> <chr> <chr> <chr>
## 1 To the Tidyverse and Beyond: Challenges fo~ Dianne Co~ <NA> <NA>
## 2 Tidy spatial data analysis Edzer Peb~ Invited <NA>
## 3 Scaling Shiny apps with async programming Joe Cheng <NA> RStudio
## 4 RStudio 1.1 new features Kevin Ush~ <NA> RStudio
## 5 The future of time series and financial an~ Davis Vau~ <NA> <NA>
## 6 Developing robust shiny apps with regressi~ Winston C~ <NA> RStudioOh no! The left join has created 2 new columns, Status.x and Status.y, representing the value of Status in the x and y.
This is because all of the RStudio staff already exist in author_tbl, with a Status value of NA.
Sad times! However, a quick trip to Stack Overflow tells me what I need to do: Note that more efficient solutions may exist!
Extract all of the speakers who are not RStudio staff from
author_tblusinganti_join().Join my
rstudiotable with my originalscheduletable which does not have aStatuscolumn, usingleft_join(). As I removed duplicates after this, I’ll have to do it again, usingdistinct().Combine the outputs from 1 and 2, using
bind_rows(). It looks kinda ugly below as I wanted to do it all in a single pipe.
I’ve used tail() to have a peek at the last 6 rows of the data_frame as I expect the rows for all the RStudio staff to be at the bottom of the table.
author_tbl <- anti_join(author_tbl, rstudio, by = "authors") %>%
bind_rows(
left_join(rstudio, schedule, by = "authors") %>%
distinct()
)
tail(author_tbl)## # A tibble: 6 x 3
## talks authors Status
## <chr> <chr> <chr>
## 1 Debugging techniques in RStudio Amanda Gadrow RStud~
## 2 Machine Learning with R and TensorFlow JJ Allaire RStud~
## 3 Building Spark ML pipelines with sparklyr Kevin Kuo RStud~
## 4 Deploying TensorFlow models with tfdeploy Javier Luras~ RStud~
## 5 Large scale machine learning using TensorFlow, Big~ Michael Quinn RStud~
## 6 R for Presidents Tareef Kawaf RStud~Hooray!
I skim through the data and realise I probably need to add a Status value for the keynote speakers. JJ Allaire has already been included in RStudio, so it’s just Dianne Cook that I need to add. I use the same technique as before:
keynotes <- data_frame(authors = "Dianne Cook",
Status = "keynote")
author_tbl <- anti_join(author_tbl, keynotes, by = "authors") %>%
bind_rows(
inner_join(keynotes, schedule, by = "authors") %>%
distinct()
)I have a quick look over the data, and remove the discussions and closing remarks from the data using filter().
author_tbl <- author_tbl %>%
filter(!talks %in% c("R in industry discussion", "Tidyverse discussion", "Closing remarks") )Finally, I filter() again to only show the rows for which the Status column contains an NA, and drop the now-redundant Status column.
contributed_talks <- filter(author_tbl, is.na(Status)) %>%
select(-Status)
contributed_talks## # A tibble: 20 x 2
## talks authors
## <chr> <chr>
## 1 The future of time series and financial analysis in ~ Davis Vaughan
## 2 infer: a package for tidy statistical inference Andrew Bray
## 3 Tidying up your network analysis with tidygraph and ~ Thomas Lin Peder~
## 4 The lesser known stars of the tidyverse Emily Robinson
## 5 Creating interactive web graphics suitable for explo~ Carson Sievert
## 6 Open-source solutions for medical marijuana Carl Ganz
## 7 Adaptive feedback for learnr tutorials Daniel Kaplan
## 8 tidycf: Turning analysis on its head by turning cash~ Emily Riederer
## 9 Branding and automating your work with R Markdown Daniel Hadley
## 10 Understanding PCA using Shiny and Stack Overflow data Julia Silge
## 11 Connecting to open source databases Kirill Müller
## 12 An assignment operator to unpack vectors and lists Nathan Teetor
## 13 Developing and deploying large scale shiny applicati~ Herman Sontrop
## 14 Five packages in five weeks - from boredom to contri~ Giora Simchoni
## 15 A SAS-to-R success story Elizabeth J. Atk~
## 16 Reinforcement learning in Minecraft with CNTK-R Ali Zaidi
## 17 Kaggle in the classroom: using R and GitHub to run p~ Colin Rundel
## 18 Imagine Boston 2030: Using R-Shiny to keep ourselves~ Kayla Patel
## 19 Something old, something new, something borrowed, so~ Chester Ismay
## 20 Training an army of new data scientists Marco BlumeI’m now left with around 20 talks, which is the number I expected. Hooray!
The final step is to get hold of the abstracts.
Getting the URLs of the Abstracts
The only place I could find the abstracts for the talks was on the individual videos, so there’s a final bit of webscraping to do.
Like before, I used read_html() to pull the entire page, and then html_nodes() and html_text() to pull out the text.
video_links <- read_html("https://www.rstudio.com/resources/videos/rstudioconf-2018-talks/")
titles <- video_links %>%
html_nodes("#post-15671 a") %>%
html_text()
# Get rid of any blank values
titles <- titles[titles!=""]
head(titles)## [1] "Greg Swinehart"
## [2] "The unreasonable effectiveness of empathy"
## [3] "Teach the Tidyverse to beginners"
## [4] "How I Learned to Stop Worrying and Love the Firewall"
## [5] "Imagine Boston 2030: Using R-Shiny to keep ourselves accountable and empower the public"
## [6] "Phrasing: communicating data science through tweets, gifs, and classic misdirection"I also need the links to the pages where the talks are. To pull these out, instead of html_text(), I used html_attr() which extracts everything with a specified attribute, in this case, “href” for the link locations.
As there is duplication I only keep every alternating value.
links <- video_links %>%
html_nodes("#post-15671 a") %>%
html_attr("href")
links <- links[seq(from = 1, to = length(links), by = 2)]
head(links)## [1] "https://www.rstudio.com/rviews/author/greg/"
## [2] "https://www.rstudio.com/resources/videos/the-unreasonable-effectiveness-of-empathy/"
## [3] "https://www.rstudio.com/resources/videos/teach-the-tidyverse-to-beginners/"
## [4] "https://www.rstudio.com/resources/videos/how-i-learned-to-stop-worrying-and-love-the-firewall/"
## [5] "https://www.rstudio.com/resources/videos/imagine-boston-2030-using-r-shiny-to-keep-ourselves-accountable-and-empower-the-public/"
## [6] "https://www.rstudio.com/resources/videos/phrasing-communicating-data-science-through-tweets-gifs-and-classic-misdirection/"I then grab the presenter names…
names <- video_links %>%
html_nodes("em") %>%
html_text()
head(names)## [1] "JD Long" "David Robinson" "Ian Lyttle" "Kayla Patel"
## [5] "Mara Averick" "Tanya Cashorali"Great! Now I’m ready to combine my titles, links, and names. If we look at the first few elements of names, titles and links, we can see that there is an erroneous extra value in titles and links but not names - I must have accidentally picked this up with my selectors from earlier.
head(names, 3)## [1] "JD Long" "David Robinson" "Ian Lyttle"head(titles, 3)## [1] "Greg Swinehart"
## [2] "The unreasonable effectiveness of empathy"
## [3] "Teach the Tidyverse to beginners"head(links, 3)## [1] "https://www.rstudio.com/rviews/author/greg/"
## [2] "https://www.rstudio.com/resources/videos/the-unreasonable-effectiveness-of-empathy/"
## [3] "https://www.rstudio.com/resources/videos/teach-the-tidyverse-to-beginners/"To get rid of this, I combine names and titles in a data_frame and then slice() this extra row off before adding the name column.
abstract_links <- data_frame(titles, links) %>%
slice(-1) %>%
mutate(name = names)
head(abstract_links)## # A tibble: 6 x 3
## titles links name
## <chr> <chr> <chr>
## 1 The unreasonable effectiveness~ https://www.rstudio.com/resour~ JD Long
## 2 Teach the Tidyverse to beginne~ https://www.rstudio.com/resour~ David R~
## 3 How I Learned to Stop Worrying~ https://www.rstudio.com/resour~ Ian Lyt~
## 4 Imagine Boston 2030: Using R-S~ https://www.rstudio.com/resour~ Kayla P~
## 5 Phrasing: communicating data s~ https://www.rstudio.com/resour~ Mara Av~
## 6 Rapid prototyping data product~ https://www.rstudio.com/resour~ Tanya C~Awesome, I have my table of URLs, talks, and authors. Let’s join that with my table of contributed talks.
left_join(contributed_talks, abstract_links, by = c("authors" = "name"))There are a couple of talks missing URLS and further inspection reveals that this is due to a missing middle name and a missing umlaut, so I make a couple of manual adjustments using case_when() and redo my join.
talks_and_urls <- contributed_talks %>%
mutate(
authors = case_when(
authors == "Ali Zaidi" ~ "Ali-Kazim Zaidi",
authors == "Kirill Müller" ~ "Kirill Muller",
TRUE ~ authors
)
) %>%
left_join(abstract_links, by = c("authors" = "name"))
talks_and_urlsAnd it’s worked! But what I want is the actual abstract text, which is what I’ll be doing in the next section.
Scraping the Abstracts
In order to scrape the abstracts from the table of talks and URLs, I need to:
- Read each URL
- Use the relevant selector to pull out just the abstract text nodes
- Use
html_text()to pull just the text out of each node.
I’ve done this a few times already, so it should be simple case of chucking this all into an lapply(), right? Well, not quite…
abstracts <- lapply(talks_and_urls$links, function(i){
read_html(i) %>%
html_nodes(".2_3 p") %>%
html_text()
})## Error in parse_simple_selector(stream): Expected selector, got <NUMBER '.2' at 1>I quickly get an error. I’m not sure of the source of this, but a remedy here is to use XPath instead of CSS (easily acquired from Selector Gadget) to specify the components I want:
abstracts <- lapply(talks_and_urls$links, function(i){
read_html(i) %>%
html_nodes(xpath='//*[contains(concat( " ", @class, " " ), concat( " ", "2_3", " " ))]//p') %>%
html_text()
})This takes a while as there are a lof of asbtracts to pull, but once it’s done, it’s looking good. This object is a list, each element containing a character vector of length 1 or more for each abstract. I want to make sure they’re all only 1 item long, so use vapply() to paste() the contents of each abstract into a single character object.
abstracts <- vapply(abstracts, function(abstract){
paste(abstract, collapse = " ")
}, character(1))Finally, I append the abstracts column onto my talks_and_urls table.
talks_and_urls$abstracts = abstractsSo here we have it, the full list of abstracts. I’m not 100% sure it’s accurate, as this post on the RStudio blog suggests different numbers of people for each status. My best guess is that couple of people I have down as delivering “contributed” talks were actually invited speakers who were invited after the publication of this blog post.